Phenotypic heterogeneity, one cell at a time
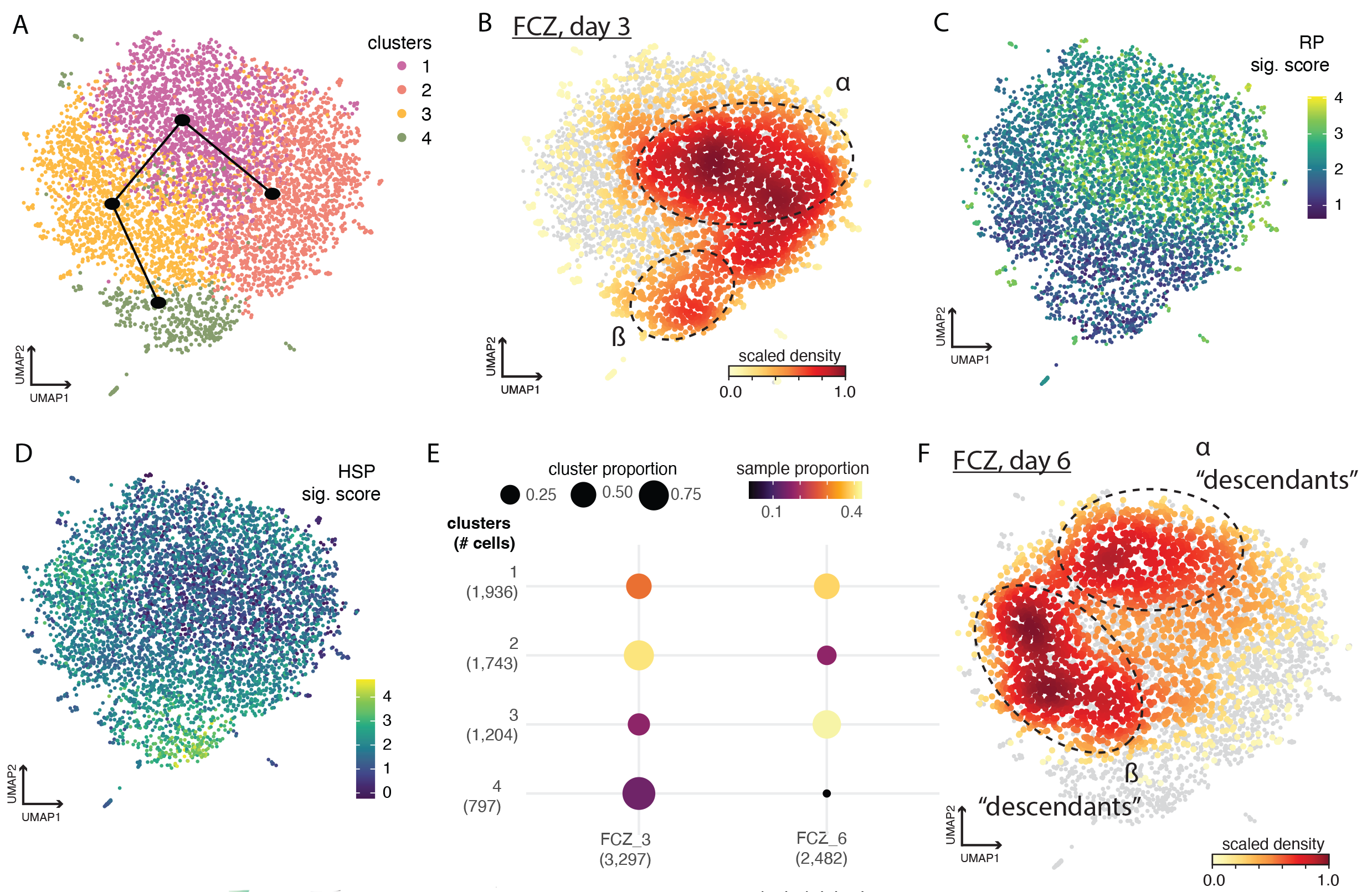
Genetically identical microbes do not all behave the same way. Within an isogenic population, individual cells diverge in transcriptional state, metabolism, and survival strategy — a phenomenon known as phenotypic heterogeneity. In pathogens like Candida albicans, this variability lets a small fraction of cells survive antifungal exposure and seed treatment failure. In the gut microbiome, the same principle plays out across species: heterogeneous subpopulations enable division of labor, bet-hedging, and resilience to environmental change. Bulk RNA-seq averages over all of this; we need single-cell resolution to see it.
Our eLife 2023 study was the first to tailor a nanolitre-droplet scRNA-seq assay to the fungal pathogen Candida albicans, then used it to dissect how genetically identical cells mount distinct cytoprotective programs against fluconazole, caspofungin, and nystatin. Building on that platform, we are extending these ideas to other adaptive states in C. albicans, including ongoing work with Lucy Xie (Stanford), and developing experimental and computational tooling for single-cell profiling of complex microbial communities, where current methods still struggle.
Computational approaches for community-scale function
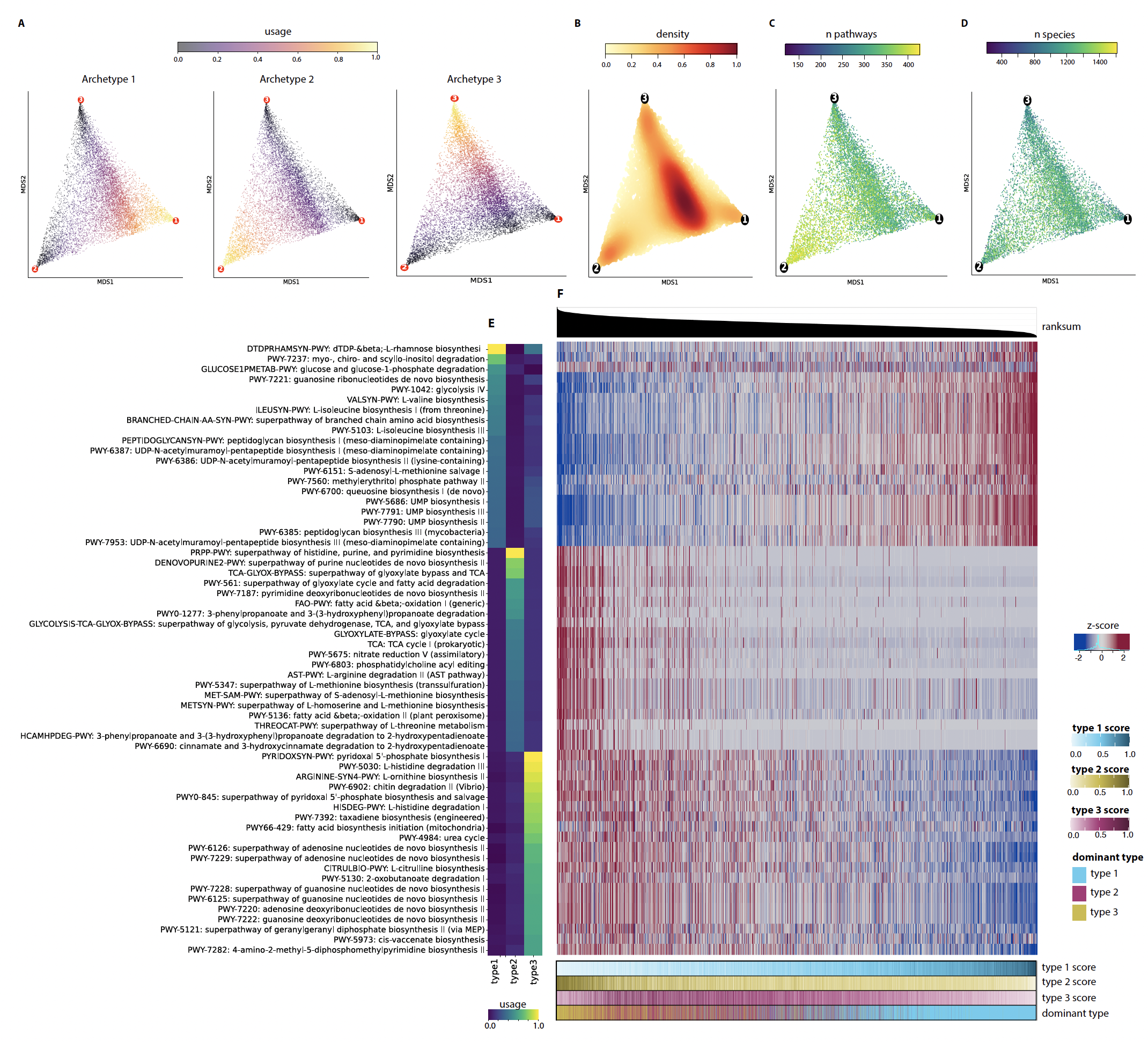
Resolving microbial behaviour cell by cell is one half of the picture; understanding how those cells organize into functional communities is the other. We develop computational approaches that integrate large-scale metagenomic data to characterize how microbes assemble, interact, and contribute to host physiology.
One example is deep-fMC, our deep-learning framework for identifying functional microbial configurations in whole-population gut metagenomes. In an analysis of nearly 10,000 healthy adult samples across 29 countries, we found that the gut’s functional space is bounded by three archetypes: one centred on sugar metabolism, one on fatty-acid metabolism, one on amino-acid and nitrogen metabolism. Most individuals are blends. Proximity to a single archetype tracks with stability, and the archetypes themselves act as confounders in disease-associated signatures across type-2 diabetes, colorectal cancer, and inflammatory bowel disease.
deep-fMC is one piece of a broader programme. We are also developing approaches to model microbe–microbe and microbe–host interactions, link community-scale functional descriptions to the cellular and sub-community dynamics that produce them, and identify the contexts in which a perturbation (diet, drug, dysbiosis) actually shifts a community’s functional state.
Completed projects
Contribution of the brain–gut–microbiota axis in health

Each human gut harbors trillions of microbes whose collective genome (the microbiome) outnumbers the host’s by 150–300×. Beyond digestion, this community shapes immune tone, metabolism, and even brain function via the brain–gut–microbiome axis: gas, chemical, and neurotransmitter signals from the gut feed back to brain regions controlling stress, satiation, sleep, memory, and cognition. Because the microbiome is itself highly variable between individuals, single-snapshot studies are easy to over-interpret. As a pilot, we profiled gut microbiota and brain activity in the same individuals before and after an exercise training intervention, pairing each subject to themselves to separate microbiome-driven effects from baseline interpersonal variation. The study is complete, and we are now using the stool samples it generated to compare bulk and single-cell metatranscriptomics, an area where methods are still maturing and where the two views can give quite different pictures of microbial activity.
Selected papers
- Functional archetypes in the human gut microbiome reveal metabolic diversity, stability, and influence disease-associated signatures
- Candida albicans exhibits heterogeneous and adaptive cytoprotective responses to anti-fungal compounds
- A metagenomic-based study of two sites from the Barbadian reef system
Selected software
- deep-fMC — deep-learning identification of functional microbial configurations.
- Candescence — automated classification of C. albicans morphologies.